Zfp36 (基因名), mRNA decay activator protein ZFP36 (蛋白名), ttp_mouse.
产品名称:
Mouse Zfp36/ mRNA decay activator protein ZFP36 ELISA Kit
Tristetraprolin蛋白
货号:
E5400m
商标:
EIAab®
监管等级:
别名:
Growth factor-inducible nuclear protein NUP475, TPA-induced sequence 11, Tristetraprolin, Zinc finger protein 36, Zfp-36, Nup475, Tis11, Tis11a, Ttp
检测方法:
ELISA
实验类型:
Sandwich
检测范围:
78-5000pg/mL
灵敏度:
37.6pg/mL
特异性:
Natural and recombinant mouse mRNA decay activator protein ZFP36
样品类型:
Serum, plasma, tissue homogenates, cell culture supernates and other biological fluids
样品数据:
登录.
实验步骤:

研究领域:
Immunology
精密度
批内差:已知浓度的3个样本在一个板子内重复检测20次,以评估批内精密度。
批内 CV: 0
批间差:已知浓度的3个样本在不同的板子上重复测定5次,以评估测定批间精密度。
批间 CV: 0
批内 CV: 0
批间差:已知浓度的3个样本在不同的板子上重复测定5次,以评估测定批间精密度。
批间 CV: 0
回收率
回收率:低、中和高浓度的分析物被掺入到血清或者血浆样本中,进行回收实验测定。
Sample Type | Average(%) | Recovery Range(%) |
Serum | 80 | 80-80 |
Plasma | 80 | 80-80 |
线性
线性:给定样本通过梯度稀释,每次稀释的测量值与理论值的比值。
Sample | 1:2 | 1:4 | 1:8 | 1:16 |
serum(n=5) | 108-120% | 110-120% | 105-115% | 104-114% |
EDTA plasma(n=5) | 106-118% | 90-100% | 108-120% | 104-113% |
heparin plasma(n=5) | 100-109%
| 106-116% | 83-94% | 108-120% |
通用注释
亚单元:
Associates with cytoplasmic CCR4-NOT and PAN2-PAN3 deadenylase complexes to trigger ARE-containing mRNA deadenylation and decay processes (PubMed:20595389, PubMed:21078877). Part of a mRNA decay activation complex at least composed of poly(A)-specific exoribonucleases CNOT6, EXOSC2 and XRN1 and mRNA-decapping enzymes DCP1A and DCP2 (By similarity). Associates with the RNA exosome complex (By similarity). Interacts (via phosphorylated form) with 14-3-3 proteins; these interactions promote exclusion of ZFP36 from cytoplasmic stress granules in response to arsenite treatment in a MAPKAPK2-dependent manner and does not prevent CCR4-NOT deadenylase complex recruitment or ZFP36-induced ARE-containing mRNA deadenylation and decay processes (PubMed:15014438, PubMed:20595389). Interacts with 14-3-3 proteins; these interactions occur in response to rapamycin in an Akt-dependent manner (By similarity). Interacts with AGO2 and AGO4 (By similarity). Interacts (via C-terminus) with CNOT1; this interaction occurs in a RNA-independent manner and induces mRNA deadenylation (PubMed:21278420). Interacts (via N-terminus) with CNOT6 (By similarity). Interacts with CNOT6L (PubMed:21078877). Interacts (via C-terminus) with CNOT7; this interaction occurs in a RNA-independent manner, induces mRNA deadenylation and is inhibited in a phosphorylation MAPKAPK2-dependent manner (PubMed:20595389, PubMed:21278420). Interacts (via unphosphorylated form) with CNOT8; this interaction occurs in a RNA-independent manner and is inhibited in a phosphorylation MAPKAPK2-dependent manner (PubMed:20595389). Interacts with DCP1A (By similarity). Interacts (via N-terminus) with DCP2 (By similarity). Interacts with EDC3 (By similarity). Interacts (via N-terminus) with EXOSC2 (By similarity). Interacts with heat shock 70 kDa proteins (By similarity). Interacts with KHSRP; this interaction increases upon cytokine-induced treatment (By similarity). Interacts with MAP3K4; this interaction enhances the association with SH3KBP1/CIN85 (By similarity). Interacts with MAPKAPK2; this interaction occurs upon skeletal muscle satellite cell activation (PubMed:25815583). Interacts with NCL (By similarity). Interacts with NUP214; this interaction increases upon lipopolysaccharide (LPS) stimulation (By similarity). Interacts with PABPC1; this interaction occurs in a RNA-dependent manner (PubMed:20595389, PubMed:21078877). Interacts (via hypophosphorylated form) with PABPN1 (via RRM domain and C-terminal arginine-rich region); this interaction occurs in the nucleus in a RNA-independent manner, decreases in presence of single-stranded poly(A) RNA-oligomer and in a p38 MAPK-dependent-manner and inhibits nuclear poly(A) tail synthesis (PubMed:22844456). Interacts with PAN2 (PubMed:21078877). Interacts (via C3H1-type zinc finger domains) with PKM (By similarity). Interacts (via C3H1-type zinc finger domains) with nuclear RNA poly(A) polymerase (PubMed:22844456). Interacts with PPP2CA; this interaction occurs in LPS-stimulated cells and induces ZFP36 dephosphorylation, and hence may promote ARE-containing mRNAs decay (PubMed:17170118). Interacts (via C-terminus) with PRR5L (via C-terminus); this interaction may accelerate ZFP36-mediated mRNA decay during stress (By similarity). Interacts (via C-terminus) with SFN; this interaction occurs in a phosphorylation-dependent manner (PubMed:11886850). Interacts (via extreme C-terminal region) with SH3KBP1/CIN85 (via SH3 domains); this interaction enhances MAP3K4-induced phosphorylation of ZFP36 at Ser-58 and Ser-85 and does not alter neither ZFP36 binding to ARE-containing transcripts nor TNF-alpha mRNA decay (By similarity). Interacts with XRN1 (By similarity). Interacts (via C-terminus and Ser-178 phosphorylated form) with YWHAB; this interaction occurs in a p38/MAPKAPK2-dependent manner, increases cytoplasmic localization of ZFP36 and protects ZFP36 from Ser-178 dephosphorylation by serine/threonine phosphatase 2A, and hence may be crucial for stabilizing ARE-containing mRNAs (PubMed:14688255, PubMed:17170118). Interacts (via phosphorylated form) with YWHAE (PubMed:21078877). Interacts (via C-terminus) with YWHAG; this interaction occurs in a phosphorylation-dependent manner (PubMed:11886850). Interacts with YWHAH; this interaction occurs in a phosphorylation-dependent manner (PubMed:11886850). Interacts with YWHAQ; this interaction occurs in a phosphorylation-dependent manner (PubMed:11886850). Interacts with (via C-terminus) YWHAZ; this interaction occurs in a phosphorylation-dependent manner (PubMed:11886850). Does not interact with SH3KBP1 (PubMed:20221403). Interacts (via the 4EHP-binding motif) with EIF4E2; the interaction is direct (By similarity). Interacts (via P-P-P-P-G repeats) with GIGYF2; the interaction is direct (PubMed:26763119).
功能:
Zinc-finger RNA-binding protein that destabilizes numerous cytoplasmic AU-rich element (ARE)-containing mRNA transcripts by promoting their poly(A) tail removal or deadenylation, and hence provide a mechanism for attenuating protein synthesis (PubMed:10330172, PubMed:10706852, PubMed:10805719, PubMed:15014438, PubMed:15187092, PubMed:15634918, PubMed:17030620, PubMed:19188452, PubMed:20595389, PubMed:21078877, PubMed:22701344, PubMed:27193233). Acts as an 3'-untranslated region (UTR) ARE mRNA-binding adapter protein to communicate signaling events to the mRNA decay machinery (PubMed:21278420). Recruits deadenylase CNOT7 (and probably the CCR4-NOT complex) via association with CNOT1, and hence promotes ARE-mediated mRNA deadenylation (PubMed:21278420). Functions also by recruiting components of the cytoplasmic RNA decay machinery to the bound ARE-containing mRNAs (PubMed:21278420). Self regulates by destabilizing its own mRNA (PubMed:15187092, PubMed:17288565). Binds to 3'-UTR ARE of numerous mRNAs and of its own mRNA (PubMed:11533235, PubMed:15187092, PubMed:16508014, PubMed:17288565, PubMed:17971298, PubMed:20595389, PubMed:21078877, PubMed:21278420, PubMed:22701344, PubMed:27193233). Plays a role in anti-inflammatory responses; suppresses tumor necrosis factor (TNF)-alpha production by stimulating ARE-mediated TNF-alpha mRNA decay and several other inflammatory ARE-containing mRNAs in interferon (IFN)- and/or lipopolysaccharide (LPS)-induced macrophages (PubMed:8630730, PubMed:9703499, PubMed:15014438, PubMed:16514065). Plays also a role in the regulation of dendritic cell maturation at the post-transcriptional level, and hence operates as part of a negative feedback loop to limit the inflammatory response (By similarity). Promotes ARE-mediated mRNA decay of hypoxia-inducible factor HIF1A mRNA during the response of endothelial cells to hypoxia (By similarity). Positively regulates early adipogenesis of preadipocytes by promoting ARE-mediated mRNA decay of immediate early genes (IEGs) (PubMed:22701344). Negatively regulates hematopoietic/erythroid cell differentiation by promoting ARE-mediated mRNA decay of the transcription factor STAT5B mRNA (By similarity). Plays a role in maintaining skeletal muscle satellite cell quiescence by promoting ARE-mediated mRNA decay of the myogenic determination factor MYOD1 mRNA (PubMed:25815583). Associates also with and regulates the expression of non-ARE-containing target mRNAs at the post-transcriptional level, such as MHC class I mRNAs (By similarity). Participates in association with argonaute RISC catalytic components in the ARE-mediated mRNA decay mechanism; assists microRNA (miRNA) targeting ARE-containing mRNAs (By similarity). May also play a role in the regulation of cytoplasmic mRNA decapping; enhances decapping of ARE-containing RNAs, in vitro (By similarity). Involved in the delivery of target ARE-mRNAs to processing bodies (PBs) (By similarity). In addition to its cytosolic mRNA-decay function, affects nuclear pre-mRNA processing (PubMed:22844456). Negatively regulates nuclear poly(A)-binding protein PABPN1-stimulated polyadenylation activity on ARE-containing pre-mRNA during LPS-stimulated macrophages (PubMed:22844456). Also involved in the regulation of stress granule (SG) and P-body (PB) formation and fusion (PubMed:15967811). Plays a role in the regulation of keratinocyte proliferation, differentiation and apoptosis (By similarity). Plays a role as a tumor suppressor by inhibiting cell proliferation in breast cancer cells.
亚细胞位置:
Nucleus
Cytoplasm
Cytoplasmic granule
Cytoplasm
P-body
Shuttles between nucleus and cytoplasm in a CRM1-dependent manner (PubMed:11796723, PubMed:11886850). Localized predominantly in the cytoplasm in a p38 MAPK- and YWHAB-dependent manner (PubMed:11886850, PubMed:16508015). Colocalizes with SH3KBP1 and MAP3K4 in the cytoplasm (By similarity). Component of cytoplasmic stress granules (SGs) (PubMed:15967811). Localizes to cytoplasmic stress granules upon energy starvation (PubMed:15014438). Localizes in processing bodies (PBs) (By similarity). Excluded from stress granules in a phosphorylation MAPKAPK2-dependent manner (PubMed:15014438). Shuttles in and out of both cytoplasmic P-body and SGs (PubMed:15967811).
该产品尚未在任何出版物中被引用。
[1].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒可以做多少个样本?
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒分为2种规格,96孔和48孔。96孔的试剂盒,标曲和样本都做复孔的话,可以检测40个样本。96孔的试剂盒,标曲和样本都不做复孔的话,可以检测88个样本。
[2].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒使用视频?
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒实验操作视频在以下网址中,对每一步的实验步骤都做了演示,方便实验员能更好地理解ELISA实验的过程。
https://www.eiaab.com.cn/lesson-tech/805.html
https://www.eiaab.com.cn/lesson-tech/805.html
[3].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒是放在-20℃冰箱保存吗?
EIAab的小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒,洗涤液、底物、终止液保存于4℃,其余试剂-20℃冰箱保存。
[4].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒原理?
双抗体夹心法:用纯化的抗体包被微孔板,制成固相抗体,往包被有固相抗体的微孔中依次加入标准品或受检样本、生物素化抗体、HRP标记的亲和素,经过彻底洗涤后用底物TMB显色。用酶标仪在450nm波长下测定吸光度(OD值),计算样本浓度。
竞争法:用纯化的抗体包被微孔板,制成固相抗体,往包被有固相抗体的微孔中依次加入标准品或受检样本和生物素标记的目标分析物,受检标本中抗原与生物素标记抗原竞争结合有限的抗体。再加入HRP标记的亲和素,经过彻底洗涤后用底物TMB显色。用酶标仪在450nm波长下测定吸光度(OD值),计算样本浓度。
竞争法:用纯化的抗体包被微孔板,制成固相抗体,往包被有固相抗体的微孔中依次加入标准品或受检样本和生物素标记的目标分析物,受检标本中抗原与生物素标记抗原竞争结合有限的抗体。再加入HRP标记的亲和素,经过彻底洗涤后用底物TMB显色。用酶标仪在450nm波长下测定吸光度(OD值),计算样本浓度。
[5].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒中需要使用的样品量是多少?
夹心法100μL/孔,竞争法50μL/孔。如样本浓度过高时,应对样本进行稀释,以使稀释后的样本符合试剂盒的检测范围,计算时再乘以相应的稀释倍数。
[6].
如何分析小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒数据?
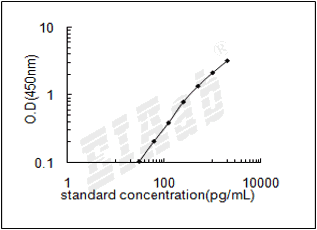
建议标准曲线,并计算样本浓度。对于elisa的曲线拟合,一般建议采用4参数曲线拟合,4参数曲线拟合通常更适合免疫分析。推荐使用专业软件进行曲线拟合,例如curve expert 1.3。根据样本的OD值由标曲查出相应的浓度,再乘以稀释倍数;或用标准物的浓度与OD值计算出标曲的回归方程式,将样本的OD值代入方程式,计算出样本浓度,再乘以稀释倍数,即为样本的实际浓度。以下链接是curve expert 1.3软件拟合曲线的方法。
https://www.eiaab.com.cn/news/502/
https://www.eiaab.com.cn/news/502/
[7].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒中是否包含人和动物的副产物,是否包含感染的或者传染性原料如HIV等?
除了抗体和稀释液中的BSA,不含其它人和动物的副产物,也不含感染材料。
[8].
收集小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒血浆样本,用什么作为抗凝剂?
一般建议用EDTA和肝素作为抗凝剂。
[9].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒酶标板可以拆成几部分?拆的时候是否需要避光,无菌?
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒酶标板是8×12孔条,可拆卸,板子可以拆成12条,注意避免孔污染,不需要避光和无菌。暂时不用的板子,放回原来装的袋子里,密封保存。
[10].
小鼠Tristetraprolin蛋白(Zfp36)ELISA试剂盒样本如何保存?
尽量检测新鲜样本。若无新鲜样本,则4℃保存1周,-20℃保存1个月,-80℃保存2个月。
反馈墙
评论数 : 0
所有用户
所有用户
默认排序
默认排序
最近
早期
目前还没有评论。



通知
规格
数量
单价 (¥)
小计 1 (¥)
小计 2:
¥

规格
数量
单价 (¥)






 验证序列:
验证序列:




 折扣:
折扣: